🦠Easier and cheaper to map resistant bacteria
A new method makes it possible to identify resistant bacteria with much simpler equipment than the ones we use today.
Share this story!
Bacteria that are resistant to antibiotics are a major problem for healthcare institutions around the world. Now there are still often opportunities to find a treatment that bites, but then we need to know which antibiotic resistance genes the bacterium carries.
To know that, we have to characterize the bacterium's DNA and it requires expensive equipment, which makes the method impractical or impossible in low-income countries. Researchers from Chalmers have now developed a method where the analysis can be done with relatively simple microscopes that are already in place in laboratories in many low-income countries.
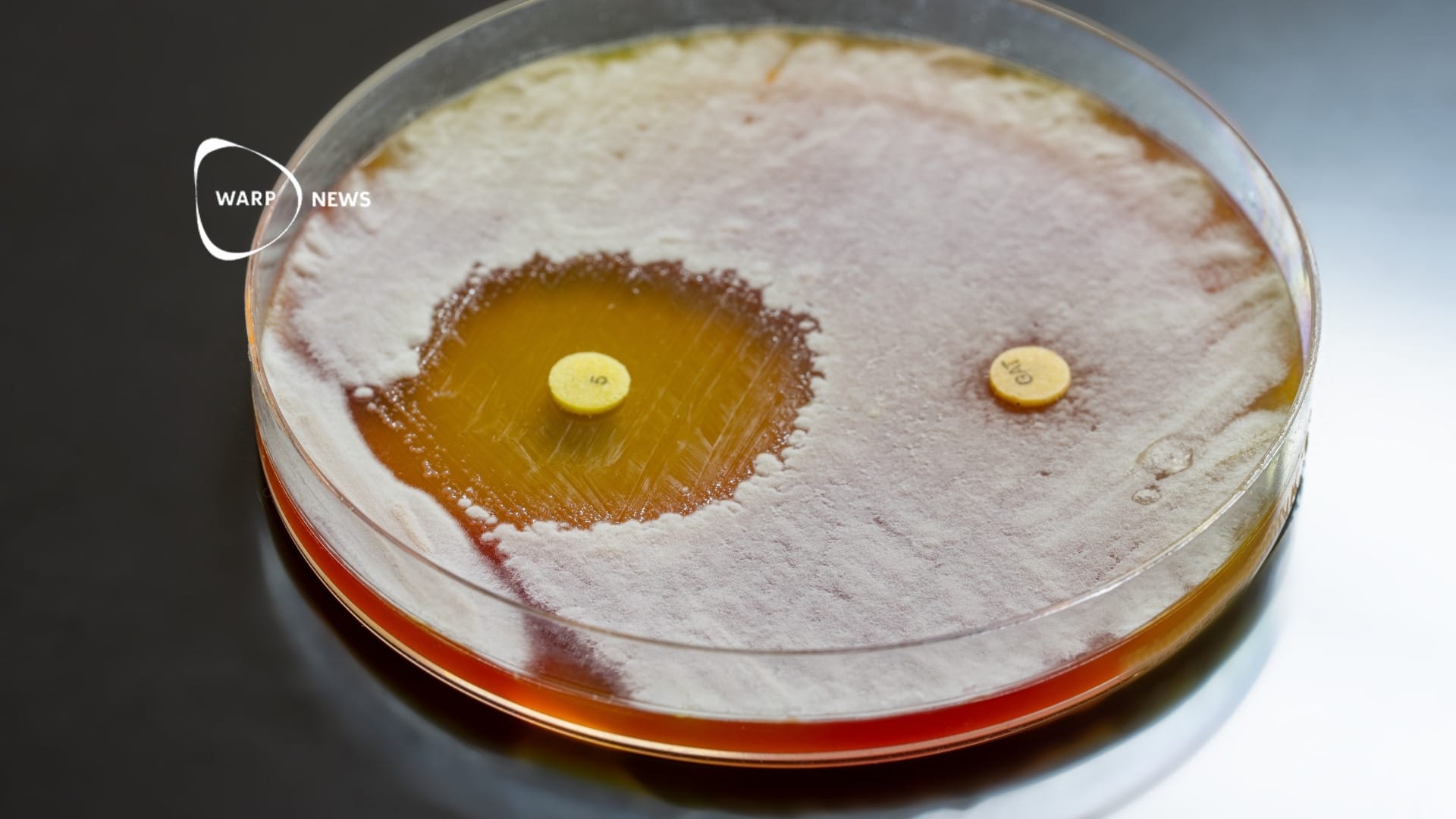
Basically, the new method is based on the Crispr-Cas9 genetic engineering technique that is being used to identify specific plasmids. Plasmids are circular DNA molecules that can show if genes in the bacterium can break down antibiotics.
- If the gene we are looking for is on the plasmid, it is cut by Cas9. When the DNA is stretched on a coverslip and imaged by fluorescence microscopy, the linear molecule can be detected. To take pictures for analysis, you can use a regular smartphone, which you easily attach to a holder by the microscope, says Gaurav Goyal, researcher at Chalmers and one of the researchers behind the study, in a press release.
To begin with, the method will be used to understand the spread of antibiotic resistance, but in the long run, the researchers hope that it can also be used to diagnose diseases that are due to resistant bacteria. The method can then also be an important tool in the richer parts of the world.
We started developing the method for less well-off laboratories, but any microbiological lab can perform this plasmid assay and get a great return on it. In addition to being able to see that a gene is present on a specific plasmid, the method can also determine the size and number of the plasmids in the sample. With our method, this is much simpler than other methods, so we also believe that it can be useful in modern microbiology labs in high-income countries, says Fredrik Westerlund, professor of chemical biology at Chalmers and another of the researchers behind the study.
By becoming a premium supporter, you help in the creation and sharing of fact-based optimistic news all over the world.